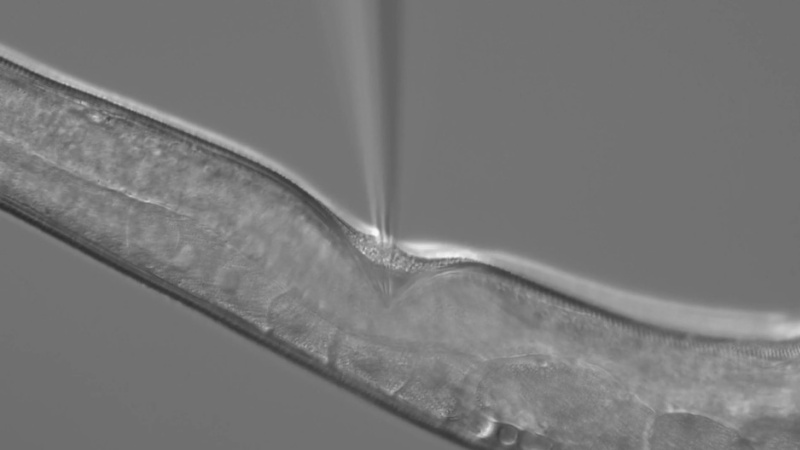
Working in teeny tiny worms, scientists can now test the effects of thousands of genetic mutations in one fell swoop.
A new gene editing technique developed by UO researchers compresses what previously would have been years of work into just a few days, making new kinds of research possible in animal models. It will allow biologists to do experiments that compare many versions of a gene, hunting for mutations that lead to specific traits and tracking their evolution over time.
Such research is often a first step towards identifying mutations that are relevant to human health or in unraveling the mechanisms behind human diseases.
While mass gene editing tricks have been developed for single-celled organisms like bacteria and yeast, this is the first time it has been possible on this scale in an animal.
“In biology, we spend a lot of time working with genetic mutants. But in animals, we're limited by how many genetic mutants we can make at one time,” said Zach Stevenson, a graduate student in the lab of biology professor and interim UO President Patrick Phillips, who helped design the technique. “This is a way around that bottleneck.”
Stevenson and his colleagues describe their new technique in preprint posted to BioRxiv.
They’ve piloted the system in C. elegans, a tiny worm that’s a popular species for biology research. A similar approach could eventually work in other lab animals, like flies or mice, Stevenson said.
“Genetic engineering the DNA of microbes has served as the basis of the revolution in biotechnology over the last three decades, but it has been difficult to do this at scale within animal systems,” Phillips said. “The new approach developed in our lab can serve as a platform for a whole new way of using a simple animal as the basis for synthetic biology in the same way that bacteria and yeast have been used for a generation."
Scientists have many reasons why they might want the ability to create many genetic mutations at once. For instance, they might be hunting for a mutation that could make an animal resistant to a specific drug, better able to survive under certain conditions, or less susceptible to a disease. They might need to screen dozens or even hundreds of possible variations on a gene to find the one that’s most effective.
Those kinds of experiments are painstakingly slow in animals. Each mutant strain, a set of worms with a specific genetic modification, must be engineered individually. Making one mutant “usually takes between seven to 10 hours of hands-on time,” Stevenson said. Using the new system, “for the same labor of making three or four mutations, you can make tens of thousands.”
To speed things up, Stevenson and his colleagues designed a way to compress hundreds or even thousands of possible mutations into a single “library.” Each book in the library is a small snippet of genetic code, meaningless and nonfunctional on its own. Each snippet fits into an engineered gap in the gene that’s being targeted, like a genetic Mad Libs puzzle.
That design means that instead of individually injecting many individual worms with different versions of a gene, researchers can inject the whole library of mutations into one worm.
Then, when the worm reproduces, the library expands.
In each offspring, one book from the mutation library gets randomly selected to complete the gene being targeted. When a segment from the gene library slides in, it makes the gene active, like flipping a switch to complete an electrical circuit.
The result: a collection of worms that all have different randomly selected genetic mutations.
The researchers named their technique TARDIS, a playful nod to the fictional Doctor Who’s space-and-time-traveling police box. Here, it stands for Transgenic Arrays Resulting in Diversity of Integrated Sequences. Like the fictional TARDIS, the worm “is bigger on the inside,” Stevenson said. That is, it contains a lot of extra genetic material.
The researchers tested TARDIS with a gene that gives worms antibiotic resistance. But they see broad applications for biology in general, including research in other model organisms.
It could be especially useful for studying interactions between proteins or signaling between cells, suggests UO research professor Stephen Banse, who helped develop TARDIS. Such interactions are often relevant for understanding disease, but scientists lose important context by studying them in yeast or bacteria, Banse said.
“Now we can do these things in an animal model,” he said.
—By Laurel Hamers, University Communications
This story was originally published by the University of Oregon on November 2, 2022.